Orbitrap Astral MS blurs the contrast between DIA and DDA
Clinical proteomics and systems biology studies require high-throughput liquid chromatography with tandem MS (LC–MS/MS) analyses with deep proteome coverage and accurate quantification. To achieve this, it is necessary to reduce MS measurement time by deploying shorter LC gradients with faster scanning MS instruments that can cope with the higher sample complexity per unit time. DIA has become the method of choice for single-shot deep proteome profiling with short gradients due to its high reproducibility and coverage and excellent quantitative performance7,11.
In DIA, co-eluting peptide ions are co-isolated and fragmented in predefined mass isolation windows, resulting in complex spectra containing fragment ions from multiple peptides. Conversely, DDA has higher specificity due to narrower isolation windows but with inferior peptide sequencing capacity. Although DIA can achieve higher specificity by constraining the quadrupole isolation width similar to DDA, state-of-the-art mass spectrometers cannot provide the sensitivity and acquisition speed needed to routinely perform DIA with 2-Th isolation windows on a chromatographic time-scale. An improved mass analyzer is required to address the trade-off between speed versus sensitivity in the Orbitrap and limited ion transmission in conventional ToF analyzers. Here, we leverage the asymmetric track lossless (Astral) mass analyzer, capable of ~200-Hz MS/MS acquisition rates at high resolving power and sensitivity, which permits routine nDIA analysis with narrow 2-Th isolation windows. The analyzer is a part of the Thermo Scientific Orbitrap Astral mass spectrometer12, which combines a modified Thermo Scientific Orbitrap Exploris 480 (ref. 13) mass spectrometer with the Astral analyzer12,14,15 via a transport octapole appended on the end of the Orbitrap Exploris ion routing multipole (IRM) (Fig. 1a, Supplementary Fig. 1 and Supplementary Note 1).
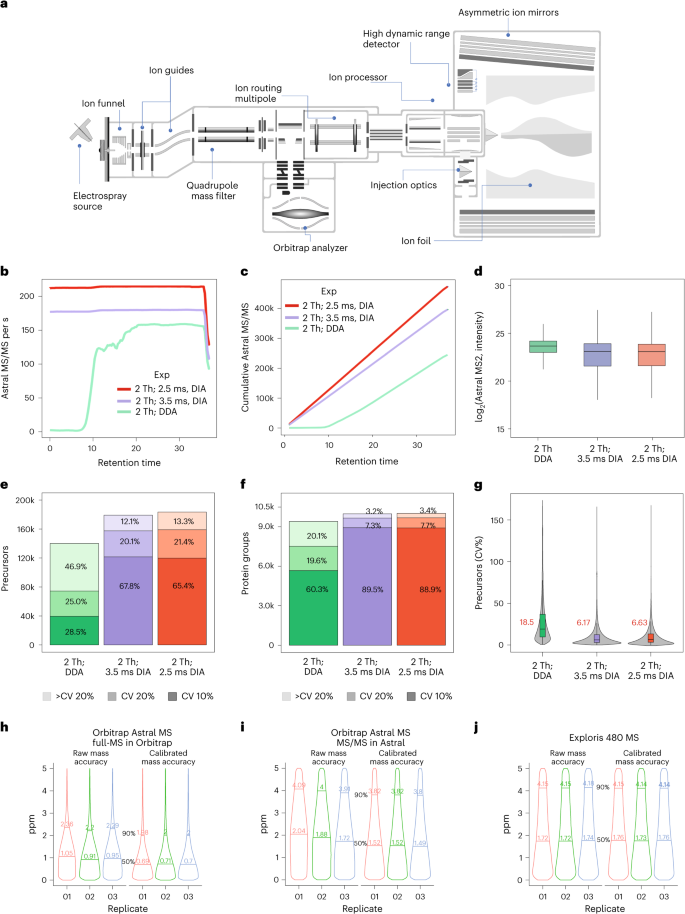
a, Hardware overview of the Orbitrap Astral MS instrument. b, MS/MS scan rate over the 28-min effective chromatographic gradient length. DDA and DIA slow (3.5 ms) and fast (2.5 ms) methods are depicted by different colors (green, purple and red, respectively). c, Cumulative MS/MS scans across the 28-min effective chromatographic gradient acquired in the Astral analyzer. d, log2 intensity of MS/MS spectra measured in the Astral analyzer with DDA and DIA. e, Quantified precursors measured with DDA and nDIA operation mode below 10% and 20% of CV (n = 3, 1,000 ng of tryptic HEK293 peptides). f, Quantified and identified proteins below 10% and 20% of CV with DDA and nDIA approaches (n = 3, 1,000 ng of tryptic HEK293 peptides). g, CVs (%) for quantified precursors by different operation modes. Median CV is shown in red for each approach. h–j, Absolute median mass deviation of peptide precursors determined with the Orbitrap Astral full-MS using the Orbitrap analyzer (h), the Orbitrap Astral MS/MS using the Astral analyzer (i) and the Orbitrap Exploris 480 MS using the Orbitrap analyzer (j); 50th and 90th quantiles are shown. In boxplot figures, center lines show the medians; box limits indicate the 25th and 75th percentiles; whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles.
Source data
In MS/MS acquisition, ions are first accumulated in the IRM, and then directed across to the ion processor, a linear quadrupole ion trap incorporating two pressure regions. The function of the ion processor is similar to the combination of a C-Trap and IRM, albeit operating much faster. Admitted ions are accumulated and fragmented via higher-energy collisional dissociation (HCD)16 in a high-pressure region, and then passed at low energy to a low-pressure region, across an apertureless interface, before final cooling and pulsed extraction to the Astral analyzer. The two regions operate in a parallelized manner such that two ion packets are processed simultaneously, in addition to a third within the IRM and any being detected with the Orbitrap and Astral analyzers, for a total of five ion packets. This approach enables the alignment of a high spectral acquisition rate with a relatively long ion processing time in the low-millisecond range. The performance envelope is particularly well suited to MS/MS operation, and in a typical analysis the Astral analyzer is set to handle all MS/MS acquisition. The coupled Orbitrap analyzer, on the other hand, is given the full length of the MS/MS cycle to acquire parallel full-MS spectra, which allows it to run at 240,000 or 480,000 resolution settings much higher than the typical 60,000 or 120,000, and which also improves full-MS sensitivity and resolution.
It is possible to operate the instrument in DDA mode with top speed acquisition of ~150 Hz, but restricting maximum ion fill time to 3.5 ms in nDIA mode results in 170-Hz MS/MS acquisition rate while injection time of maximum 2.5 ms delivers >200-Hz MS/MS acquisition (Fig. 1b and Supplementary Fig. 2a,b). The fastest nDIA scanning speed generates ~400,000 DIA MS/MS spectra in half-an-hour of LC–MS/MS analysis of a human embryonic kidney cell (HEK293) tryptic digest (Fig. 1c). The overall MS/MS spectra intensity was higher in DDA mode (Fig. 1d), reflecting the abundance bias of DDA methods having the propensity to isolate the most abundant precursors17 (Supplementary Fig. 2c). Despite lower MS/MS intensities, nDIA acquisition mode resulted in higher identifications of approximately 170,000 peptide precursors and ~10,000 protein groups (hereafter referred to as proteins) (Fig. 1e,f). The quantitation of peptides and protein identifications can be performed with high reproducibility, with coefficient of variation (CV) below 20% for 90% of precursors and 95% of proteins, respectively (Fig. 1e,f). The median CVs at precursor level for nDIA were lower compared with DDA (<7% versus <19%, respectively) (Fig. 1g). This observation can likely be explained by the semi-stochastic nature of DDA precursor selection18. The comparison between DDA and DIA strategies showed higher performance for nDIA, although closely followed by DDA and also with faster LC gradients and different DDA acquisition schemes with 15-min LC–MS/MS runs (Supplementary Fig. 2d–f). Of the four DDA acquisition methods compared, three of them, top 100, top 75 and cycle 0.5 s, showed comparable performance, with >70,000 quantified precursors in 15 min at the top speed acquisition of ~150 Hz. However, DDA cycle 0.5 s and top 75 showed the best protein identifications, with 7,378 and 7,061 quantified proteins, respectively. Despite that the DIA-NN software was originally designed for DIA data, it is superior to most DDA-specific search engines for analyzing Orbitrap Astral DDA data. We therefore used DIA-NN for analysis of both DIA and DDA data to maintain the same bioinformatics workflow for both methods.
To ensure reliable identifications from the large number of DIA MS/MS scans recorded with the Astral analyzer, we estimated the empirical false discovery rate (FDR) using entrapment database analyses19 with spectral library-free searches in Spectronaut (v.18). First, we searched against a 9× larger shuffled human mimic database. The empirical FDR at protein level corrected for the mimic database size was estimated to be ~0.7% when analyzing both HEK293 and HeLa datasets (Supplementary Fig. 3a,b). Second, we also estimated empirical FDR by searching against an Arabidopsis thaliana database as decoy and performed FDR estimations by conducting bootstrap analyses with 23 repetitions. Reassuringly, the corrected FDR estimation from this was 1.25%, consistent with the mimic database entrapment analysis (Supplementary Fig. 3c,d). Furthermore, by using Spectronaut’s algorithm for interference detection and analyzing all precursors identified, we found that only 1.4% had interfering ions (Supplementary Fig. 3e,f).
Mass accuracy determines fragment ion identity in peptide search engines20. To evaluate the Astral mass analyzer’s accuracy in MS/MS mode, we used a 200-Hz, 2-Th nDIA analysis of a tryptic digest of HEK293 cell lysate. The Orbitrap MS1 full scan at 240,000 resolution showed 0.95-ppm mass median absolute deviation (Fig. 1h), akin to previous reports21. For matched fragment ions recorded by the Astral analyzer, the median absolute deviation was 1.88 ppm, similar to Orbitrap MS/MS at 15,000 resolution (Fig. 1i,j). Although the Orbitrap mass calibration remains stable for days, the Astral mass accuracy may drift (~3 ppm) over time. This drift can be corrected through regular automatic recalibration via the internal calibrant source or post-acquisition recalibration, resulting in sub-ppm-level mass accuracy.
Fast sequencing enables deep proteome profiling
Comprehensive proteome coverage is essential for large-scale systems biology studies involving thousands of conditions. The first complete proteome of a eukaryotic organism analyzed by MS was that of yeast22, and with state-of-the-art instrumentation it was possible to cover ~4,000 yeast proteins in approximately 1 h of LC–MS/MS time23.
Analyzing whole yeast cell digests with nDIA (2-Th) across various LC gradient lengths, from 5-min active LC gradients (180 SPD) up to 96 SPD (15-min gradient), achieved nearly complete yeast proteome coverage (~4,500 proteins) ten times faster than previous reports, and maintaining high quantitative reproducibility (Fig. 2a).
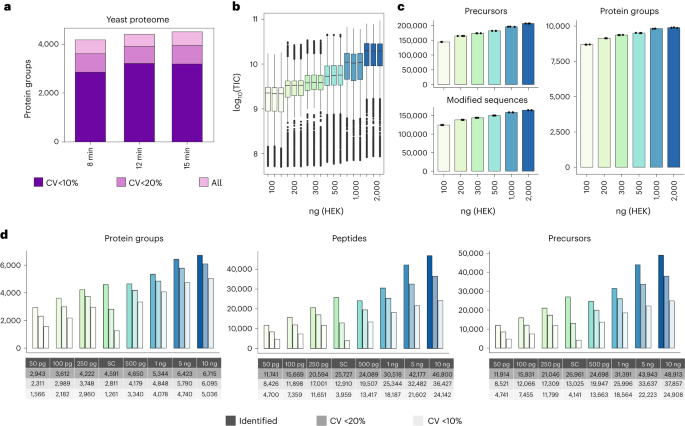
a, Protein groups identified and quantified below 10% and 20% of CV (n = 3) in a yeast extract using different chromatographic gradient lengths. b, Boxplot of total ion current (in log10 scale) measured in a dilution series (from 100 to 2,000 ng) of a HEK293 cell protein digest. c, Protein groups, precursors and modified peptides identified (n = 3) in a dilution series of HEK293 cell peptide extract using Spectronaut (v.17). d, Protein groups, peptides and precursors identified in three replicates in a low-amount dilution series and 12 single HeLa cells (SC) individually using Spectronaut (v.18). Next to each bar, the number of each quantified with CV < 20% and CV < 10%. Box limits indicate the 25th and 75th percentiles as determined by R software; whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles; outliers are represented by dots. TIC, total ion chromatogram.
Source data
In contrast, a human cell proteome comprises >12,000 (refs. 3,4,24) protein-coding genes with a high dynamic range (>6-logs)25,26, demanding extended MS measurement time for comprehensive coverage. To evaluate human proteome coverage and sensitivity achieved with nDIA (2-Th), we measured increasing amounts of HEK293 digests (from 100 ng to 2,000 ng) with 60-min active LC gradients. Reassuringly, the measured MS/MS signal scaled with loading amounts and did not seem to be saturated at the highest load (Fig. 2b). Peptide and protein coverage and quantitative reproducibility also correlated with loading amounts, with an average of 9,879 proteins and 159,627 unique peptides (164,216 modified peptides) when injecting 2 µg of HEK293 protein digest, but covering 88% of proteins with only 100 ng (Fig. 2c and Supplementary Table 1). To optimize proteome coverage and MS measurement time, we compared different loadings of the HEK293 digest across different LC gradient lengths (Supplementary Table 2, Extended Data Fig. 1 and Supplementary Note 2). This analysis revealed that the 28-min active LC gradient of 1 µg resulted in 9,619 proteins with outstanding reproducibility (median CV < 3%) (Supplementary Table 3).
Reduced identifications with lower sample loads highlight the necessity for tailored DIA acquisition methods, especially for high-sensitivity strategies. To address this, we widened the isolation window and scaled the maximum injection time (maxIT) for decreasing sample amounts, while maintaining a constant scan cycle time (Supplementary Fig. 4a). We analyzed 10-ng, 25-ng and 100-ng injections using 2-Th, 4-Th, 8-Th and 16-Th windows, with 3.5-ms, 7-ms, 14-ms and 28-ms maxITs (Supplementary Table 2), respectively, using a 60-min active LC gradient. We found that for 100 ng, the optimal method was 4-Th windows with 7-ms maxIT, yielding 8,600 proteins. Lower loads benefited from wider 8-Th windows, resulting in 6,600 proteins from just 10-ng input (Supplementary Fig. 4b and Supplementary Table 4). Decreasing loading amounts rarely reached the ideal ion target value and this under-sampling is most pronounced with narrow isolation windows (Supplementary Fig. 4c). The impact of different window sizes was not uniform across the mass range, with peptides of higher m/z most affected (Supplementary Fig. 4d). Therefore, scaling window size and maxIT in different mass ranges according to sample loads may provide a means for maximizing peptide coverage given the sample input. To test the instrument sensitivity, we performed DIA analyses on a range of HeLa digest dilutions, spanning from 50 pg up to 10 ng, and also included analysis of 12 individual single HeLa cell preparations (Fig. 2d). To maximize sensitivity of the MS/MS scans, the DIA isolation windows were gradually widened from 5 Th to 20 Th, scaling maxIT accordingly, and using high-Field Asymmetric waveform Ion Mobility Spectrometry (FAIMS) with single compensation voltage of −50 V optimized for DIA-based proteome analyses 13. The reason for adopting this approach was to improve coverage of peptides with lower abundance, otherwise missed with narrower isolation windows, while FAIMS acted as an ion filter to minimize singly charged background ions, thereby enhancing multiply charged peptide signal-to-noise ratios. As expected, peptide and protein coverage scaled with input amount, but reproducible quantification was maintained even in the low-abundance range (Fig. 2d). These data establish the high sensitivity of the Orbitrap Astral MS instrument, enabling the identification of ~3,000 proteins from 50 pg and >4,500 proteins from single HeLa cells. Approximately 80% of these proteins were quantified with CV < 20% from 250 pg in dilution series. The single-cell data exhibited higher variation, which is expected due to the inherent heterogeneity of individual cells. This dataset demonstrates that the Orbitrap Astral MS instrument is particularly well suited for single-cell proteomics research.
To benchmark the performance of the nDIA strategy on the Orbitrap Astral for high-throughput proteomics against the current state-of-the-art, we compared the 5-min HeLa analysis results with reference DIA datasets on HeLa recorded with an Orbitrap Exploris 480 (ref. 13), a ZenoTOF 7600 (ref. 27), a TripleTOF 6600 (ref. 8) or a timsTOF HT (ref. 28), also with 5-min LC gradients, respectively. When analyzed with the same DIA processing software tools (Spectronaut and DIA-NN) and software settings rendering the results comparable, the Orbitrap Astral MS instrument identified 7,538 protein groups using DIA-NN in spectral library-free mode, whereas the TripleTOF identified 3,330 proteins, the ZenoTOF 3,419 proteins, the timsTOF HT 3,737 proteins and the Orbitrap Exploris identified 3,143 proteins (Supplementary Fig. 5a). The nDIA analysis with the Orbitrap Astral reproducibly identified >75,000 different modified peptide variants with the 5-min gradient, which was 3× more identifications compared with any of the other MS platforms (Supplementary Fig. 5b).
nDIA enables precise and accurate label-free quantification
To test the nDIA method performance for label-free quantification (LFQ), we analyzed mixed species proteome samples composed of human, yeast and Escherichia coli proteins in six different ratios (Fig. 3a). We analyzed the samples using nDIA (Supplementary Table 2) to benchmark the proteome coverage and quantitative accuracy and precision of the Orbitrap Astral mass spectrometer against DIA on an Orbitrap Exploris 480 mass spectrometer.
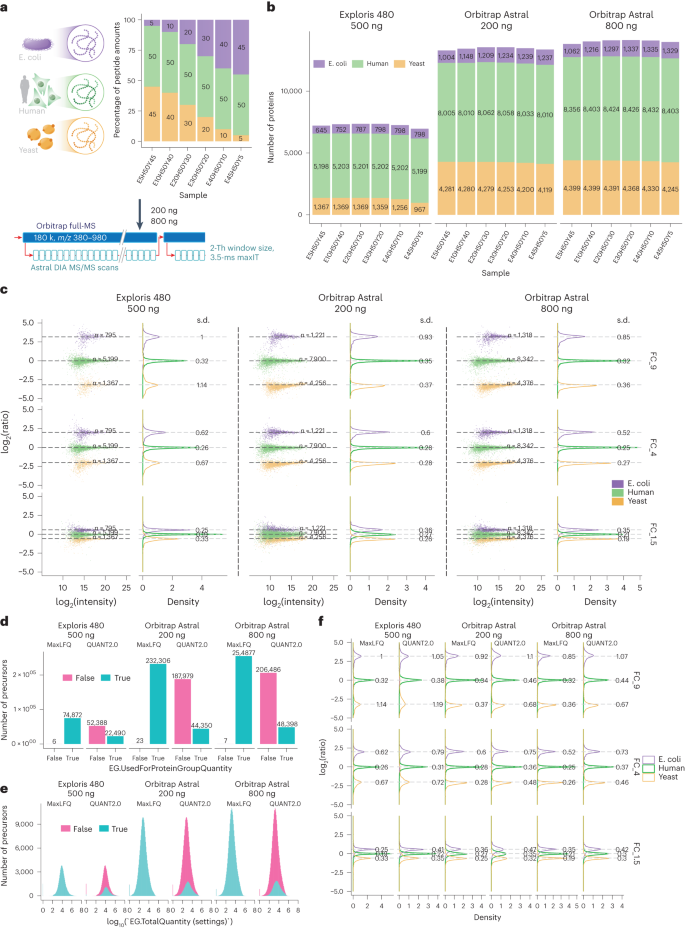
a, Graphical representation of the experimental design. Tryptic peptides from three species were combined in six distinct ratios (E5H50Y45, E10H50Y40, E20H50Y30, E30H50Y20, E40H50Y10 and E45H50Y5). Samples were processed using the Orbitrap Astral mass spectrometer in technical triplicates, employing a 3.5-ms maxIT and 2-Th window size method. The loading amounts were 200 ng and 800 ng. b, Number of proteins identified from the three species in each sample. For the Orbitrap Exploris 480 MS runs, the loading amount was 500 ng. c, log-transformed ratios of quantified proteins. Scatter plots for all runs over the log-transformed protein intensities are displayed on the left, while density plots are on the right. Colored dashed lines represent expected log2(A/B) values for proteins from humans (green), yeast (orange) and E. coli (purple). Standard deviations are displayed on the density plots. FC_9, FC_4 and FC_1.5 are calculated from log2(E45H50Y5/E5H50Y45), log2(E40H50Y10/E10H50Y40) and log2(E30H50Y20/E20H50Y30), respectively. d, Number of precursors used and not used for protein quantifications in the MaxLFQ and QUANT2.0 algorithms. e, The intensity distribution of precursors used for protein quantifications in MaxLFQ and QUANT2.0 algorithms. f, Density plots of log-transformed protein ratios quantified using both the MaxLFQ and QUANT2.0 algorithms. FC, fold change. The unit in the density plots represents a proportion of the total distribution with a sum of 100 and is normalized such that the total sums up to 100 (%).
Source data
The Orbitrap Astral MS instrument identified >2×proteins and 3.5×peptide precursors in a 28-min LC gradient when analyzing 800 ng of HEK lysate compared with the Orbitrap Exploris 480 MS instrument with a 45-min LC gradient (Fig. 3b and Supplementary Fig. 6a). Notably, the Orbitrap Astral MS instrument demonstrated high technical reproducibility, with approximately 90% of proteins having CVs < 20% and low number (~3%) of missing values in technical triplicates (Supplementary Fig. 6b,c). Additionally, the MS/MS-based protein quantitation results in the Astral analyzer showed higher accuracy and lower standard deviations (~0.3) compared with the Orbitrap Exploris 480 MS instrument at the expected ratios for yeast and E. coli (Fig. 3c). Moreover, the higher protein and peptide coverage per protein resulted in improved protein quantification, particularly when using the MaxLFQ algorithm29. Unlike the QUANT2.0 algorithm, which only uses the top three peptide groups for protein quantification, the MaxLFQ algorithm utilizes over 99% of the measured precursors for protein quantification (Fig. 3d). The QUANT2.0 algorithm disregarded ~210,000 precursors, which were almost as abundant as the ~44,000 selected30 (Fig. 3e). Including these extra precursors in the MaxLFQ algorithm significantly improved quantification performance and demonstrates that nDIA provides accurate and precise protein quantification. To asses any possible abundance bias in quantitative precision and fold accuracy, we plotted the log2-tranformed protein intensities against the corresponding CVs of the protein ratios (Supplementary Fig. 7). Reassuringly, low-abundance proteins had higher fold-change CVs compared with high-intensity proteins. Density plots depicting fold-change CV distributions per species showed that approximately ~80% of the quantified proteins for Homo sapiens and Saccharomyces cerevisiae resulted in median CV below 10%, while for E. coli it was below 20%.
Deep proteome acquisition through fast multi-shot strategy
To overcome the inherent wide dynamic range of human protein abundances, a multi-shot proteomics strategy based on offline peptide fractionation can be employed to increase dynamic range and coverage compared with single-shot experiments. Fractionation decreases sample complexity but requires extensive MS measurement time. To maximize proteome coverage, we utilized high-resolution offline HpH reversed-phase peptide chromatography31,32 in combination with short online LC gradients and nDIA (2 Th) analysis on the Orbitrap Astral MS instrument. To identify the most effective strategy to comprehensively cover a human cell line proteome, we fractionated a tryptic digest of HEK293 cells into 46, 34, 23 or 12 HpH fractions (Fig. 4a).
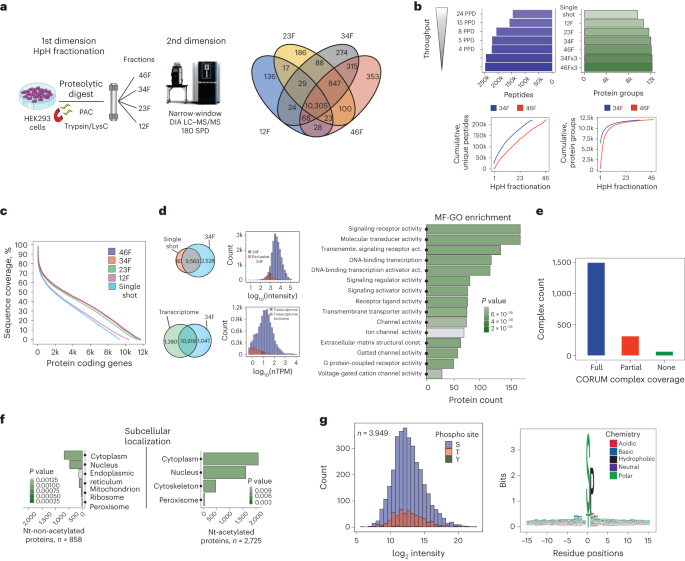
a, Experimental workflow of all HEK293 experiments and fractionation schemes (left). b, Protein and peptide identifications for different fractionation schemes and throughput are shown (top). Cumulative peptide identifications by fraction (bottom, left) and cumulative proteins (bottom, right) for both 46 and 34 fractionation schemes are presented. c, Comparison of sequence coverage achieved by different fractionation schemes and single-shot analysis. Overlap of protein groups identified between 34 fractionation scheme (5 proteomes per day (PPD)) and single-shot analysis (2-µg input and 30-min effective chromatographic gradient length). d, Protein group abundances in HEK293 identified in this dataset compared against single-shot analysis (left) are presented. GSEA functional enrichment analysis of the exclusively detected proteins in 34 fractionation scheme compared with single-shot analysis using gene ontology (GO) molecular function gene set. e, CORUM protein complex coverage of proteins identified in this dataset. f, Comparison of the GSEA functional enrichment analysis using GO cellular component terms gene set of Nt-acetylated and Nt-non-acetylated proteome. g, Abundance and sequence logo plot of detected phosphorylation sites without enrichment are shown. F, fraction; MF-GO, molecular function-gene ontology; Nt, N-terminal; nTPM, normalized transcripts per million.
Source data
By analyzing 200 ng from each of the 46 fractions with 5-min effective gradient DIA runs (6-h measurement time), we identified 12,179 proteins and 222,389 peptide sequences using Spectronaut (v.17) with directDIA+. This approach reduced the required MS acquisition time by sixfold and sample input by fivefold compared with previous approaches (Supplementary Fig. 8a). The quantitative performance in terms of reproducibility between different fractionation schemes was high, with Pearson correlation coefficients above 0.9 for all pairwise comparisons (Supplementary Fig. 8b). Additionally, ~90% of all quantified proteins in 23 HpH-fractionated HeLa and HEK cell lines had CVs < 10% (Supplementary Fig. 8c). Combining three biological replicates of each of the fractionation schemes resulted in a maximum coverage of 12,328 proteins from ~246,000 peptides from both 34 and 46 fractions (Fig. 4b). Notably, the 34 and 46 fractionation schemes achieved an average protein sequence coverage of ~40%.
Reducing the HpH fractionation scheme from 46 to 34 and 23 concatenated fractions resulted in similar proteome coverage of 99.4% and 96.7%, respectively (Fig. 4c). This suggests that routine acquisition of up to eight comprehensive human proteomes per day is feasible. The additional proteins identified by multi-shot compared with single-shot analysis are in the low-abundance range and represent important signaling proteins, including transmembrane receptors and transcription factors (Fig. 4d). The number of proteins quantified in the 34 fractionation scheme was comparable in coverage to next-generation RNA sequencing data of HEK293 (normalized transcripts per million > 0.5, 12,299 different gene products), with an overlap of 88.8% at gene level (Fig. 4d).
Our dataset encompasses ~80% of core protein complexes in the CORUM database33,34, providing evidence of its completeness (Fig. 4e). The extensive peptide coverage also enabled us to search for post-translational modifications, which typically require specific enrichment strategies before MS analysis35. We identified ~2,700 N-acetylation sites, which, as previously shown4, mainly target cytoplasmic and nuclear proteins (Fig. 4f). Additionally, we detected ~4,000 phosphorylation sites (site localization > 0.75; Supplementary Fig. 8d), covering the entire protein abundance distribution and likely representing the most abundant cellular sites (Fig. 4g). In accordance with previous studies36,37,38, phosphosites were mainly targets of proline-directed kinases such as the cyclin-dependent kinases and the mitogen-activated protein kinases.
nDIA empowers systems biology and clinical proteomics
Functional genomics screens require large-scale approaches to evaluate gene functions, often through knockouts or knockdowns39. High-throughput proteomics can be used to analyze specific biological processes or diseases caused by genetic perturbations. Historically, the model organism to perform genome-wide screens in is S. cerevisiae, and consequently the proteomics community constantly strives to acquire comprehensive yeast proteomes in minimal MS analysis time (Fig. 5a). Given the essentially complete yeast proteome profiling provided by nDIA with 5-min gradients, we applied this to analyze a yeast strain library consisting of 104 gene knockouts targeting cell cycle, proteasome, DNA damage response and kinase genes (Fig. 5b). We consistently quantified ~4,500 proteins across all biological conditions and replicates, with 93.98% of proteins reproducibly quantified with CV < 10% (Fig. 5c).
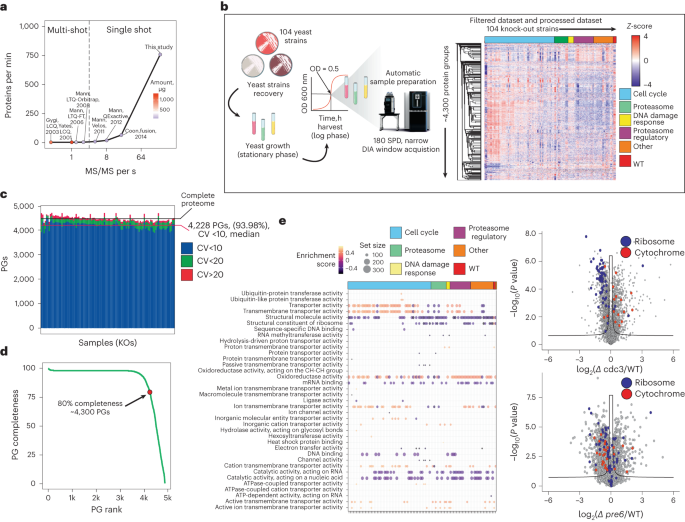
a, Rate of protein identifications as a function of mass spectrometer MS/MS scan rate for large-scale yeast proteome analysis (Yates, 2001 (ref. 53); Gygi, 2003 (ref. 52); Mann, 2006 (ref. 35); Mann, 2008 (ref. 22); Mann, 2011 (ref. 54); Mann, 2012 (ref. 55); Coon, 2014 (ref. 23)). b, Experimental setup for high-throughput yeast proteome MS analysis and dataset description. c, Protein identification numbers (mean) and CV per sample (n = 3). d, Data completeness curve showing the number of proteins quantified in a given percentage of samples. e, GSEA functional enrichment analysis results using GO molecular function terms gene set. Size of the dot indicates the set size and the color the normalized enrichment score (NES) (left). Volcano plot of differentially expressed proteins between the selected knockouts and a reference strain (right). P values were obtained by two-sided t-test and BH FDR corrected. BH, Benjamini–Hochberg; KO, knockout ; PG, protein group; WT, wild type.
Source data
In total, ~4,300 yeast proteins were consistently measured in 80% of samples (Fig. 5d). Analysis of knockout-regulated proteins showed that changes in protein abundances are linked to general biological processes such as cellular adaptation of translation rate and metabolic pathways (Fig. 5e). This dataset, collected in ~41.5 h and with great proteome depth for each strain, consistently reproduced the findings of a recent systematic yeast gene knockout screen40.
To demonstrate that nDIA is capable of achieving the high-throughput required for analyzing large clinical cohorts and conducting biomarker discovery experiments, we reanalyzed human brain protein extracts from Broadmann’s area 9 (BA9) from a multiple system atrophy (MSA) sample cohort of biopsies (n = 45 for MSA cases, and n = 29 for controls) in three technical replicates (222 LC–MS/MS runs) using the 180-SPD method (Fig. 6a). MSA is a rare and fatal neurodegenerative disease that affects most of the cortical brain areas41. Although the pathologic hallmark of MSA is accumulation of aggregated α-synuclein, the underlying pathophysiology of the disease explaining the origin of α-synuclein accumulation and the resulting loss of neurons remains unknown42. This fast nDIA analysis resulted in median coverage of 6,640 protein groups per brain sample, including more than 5,400 proteins quantified in >90% of samples. In contrast, analysis with half-an-hour active LC gradients using the Evosep One Whisper flow 40-SPD method enabled identification of 9,236 protein groups with a median of 8,320 proteins identified per sample and 7,239 proteins quantified in >90% of all samples (Fig. 6b). This comprehensive characterization of the MSA proteome was accomplished >20× faster with the 180-SPD and >5× faster with the 40-SPD method, resulting in 60% and 90% more proteins reproducibly quantified in at least 70% of the samples, respectively, compared with the original study43 (Supplementary Fig. 9a,b). The 180-SPD method quantified about 80% of the proteins quantified by 40 SPD (Fig. 6c). We were able to identify the same outliers as found originally, confirming that the status as an outlier is associated with the sample and not with the analysis method (Supplementary Fig. 9c). The nDIA analysis enabled us to reproduce the main biological findings reported in the original work, where proteins involved in blood coagulation and in the immune system (immunoglobulins and complement factors) were found with increased levels in the MSA brain parenchyma (Fig. 6d,e). The deeper proteome coverage with the 40-SPD method enabled the detection of proteins related to the glutamate release cycle and transcriptional processes, and these were highlighted through a gene set enrichment analysis (GSEA) to be elevated in the brain biopsies from patients with MSA (Fig. 6e). Altered glutamate metabolism has been proposed as a potential cause of MSA in animal models44, while disruption of transcription factor regulation has been related to the development of Parkinson-like symptoms45,46 (Supplementary Table 5). These results showcase the potential of nDIA to help identify biomarkers of biomedical importance with high throughput.
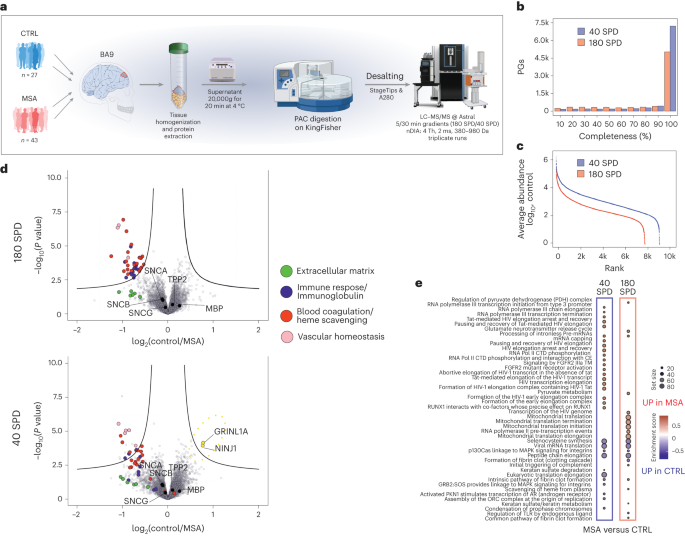
a, Sample preparation workflow of Broadmann’s area 9 (BA9) biopsies excised from either normal or MSA patient brains for proteome MS analysis. b, Histogram of data completeness showing the number of proteins quantified in a given percentage of samples for 180-SPD and 40-SPD methods. c, log10 abundance dynamic range detected by 180-SPD and 40-SPD methods. d, Volcano plots showing differential expression results between control and case samples (P values were obtained by two-sided t-test and BH FDR corrected) for both methods. e, GSEA functional enrichment analysis results using molecular pathways from the Reactome database. The top and bottom ten enriched molecular pathways for each method are shown. Size of the dot indicates the set size and the color the NES. PAC, protein aggregation capture.
Source data
Source link